Confusion matrix¶
Import ConfusionMatrix
from pandas_ml import ConfusionMatrix
Define actual values (y_true) and predicted values (y_pred)
y_true = ['rabbit', 'cat', 'rabbit', 'rabbit', 'cat', 'dog', 'dog', 'rabbit', 'rabbit', 'cat', 'dog', 'rabbit']
y_pred = ['cat', 'cat', 'rabbit', 'dog', 'cat', 'rabbit', 'dog', 'cat', 'rabbit', 'cat', 'rabbit', 'rabbit']
Let’s define a (non binary) confusion matrix
confusion_matrix = ConfusionMatrix(y_true, y_pred)
print("Confusion matrix:\n%s" % confusion_matrix)
You can see it
Predicted cat dog rabbit __all__
Actual
cat 3 0 0 3
dog 0 1 2 3
rabbit 2 1 3 6
__all__ 5 2 5 12
Matplotlib plot of a confusion matrix¶
Inside a IPython notebook add this line as first cell
%matplotlib inline
You can plot confusion matrix using:
import matplotlib.pyplot as plt
confusion_matrix.plot()
If you are not using inline mode, you need to use to show confusion matrix plot.
plt.show()
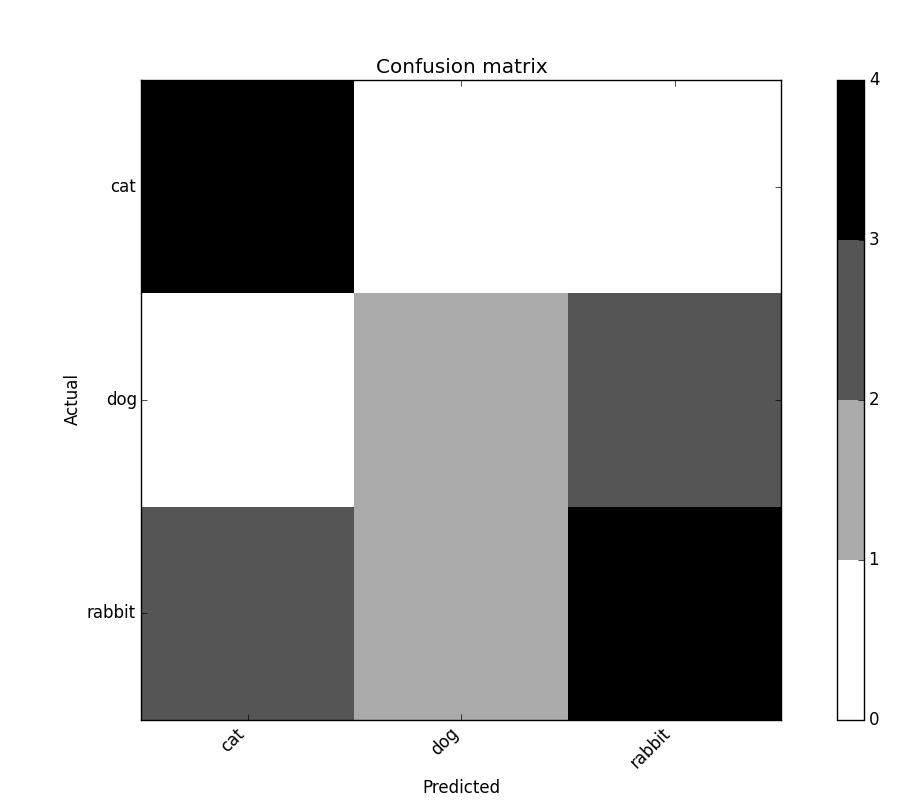
confusion_matrix
Matplotlib plot of a normalized confusion matrix¶
confusion_matrix.plot(normalized=True)
plt.show()
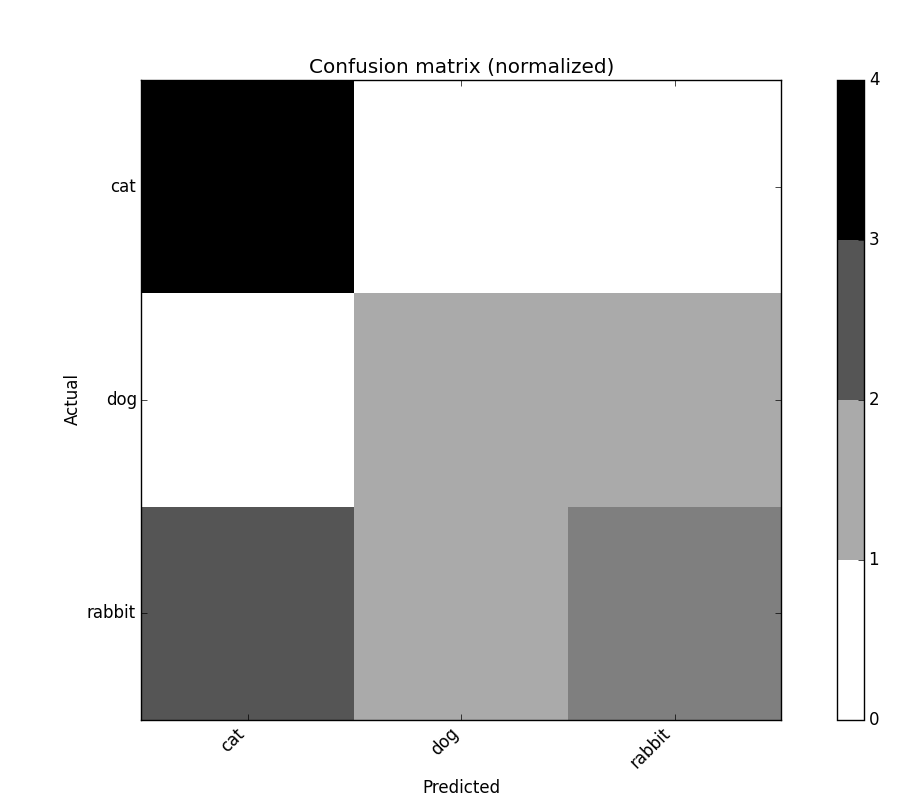
confusion_matrix_norm
Binary confusion matrix¶
If actual values (y_true) and predicted values (y_pred) are bool,
ConfusionMatrix outputs binary confusion matrix.
y_true = [ True, True, False, False, False, True, False, True, True,
False, True, False, False, False, False, False, True, False,
True, True, True, True, False, False, False, True, False,
True, False, False, False, False, True, True, False, False,
False, True, True, True, True, False, False, False, False,
True, False, False, False, False, False, False, False, False,
False, True, True, False, True, False, True, True, True,
False, False, True, False, True, False, False, True, False,
False, False, False, False, False, False, False, True, False,
True, True, True, True, False, False, True, False, True,
True, False, True, False, True, False, False, True, True,
False, False, True, True, False, False, False, False, False,
False, True, True, False]
y_pred = [False, False, False, False, False, True, False, False, True,
False, True, False, False, False, False, False, False, False,
True, True, True, True, False, False, False, False, False,
False, False, False, False, False, True, False, False, False,
False, True, False, False, False, False, False, False, False,
True, False, False, False, False, False, False, False, False,
False, True, False, False, False, False, False, False, False,
False, False, True, False, False, False, False, True, False,
False, False, False, False, False, False, False, True, False,
False, True, False, False, False, False, True, False, True,
True, False, False, False, True, False, False, True, True,
False, False, True, True, False, False, False, False, False,
False, True, False, False]
binary_confusion_matrix = ConfusionMatrix(y_true, y_pred)
print("Binary confusion matrix:\n%s" % binary_confusion_matrix)
It display as a nicely labeled Pandas DataFrame
Binary confusion matrix:
Predicted False True __all__
Actual
False 67 0 67
True 21 24 45
__all__ 88 24 112
You can get useful attributes such as True Positive (TP), True Negative (TN) …
print(binary_confusion_matrix.TP)
Matplotlib plot of a binary confusion matrix¶
binary_confusion_matrix.plot()
plt.show()
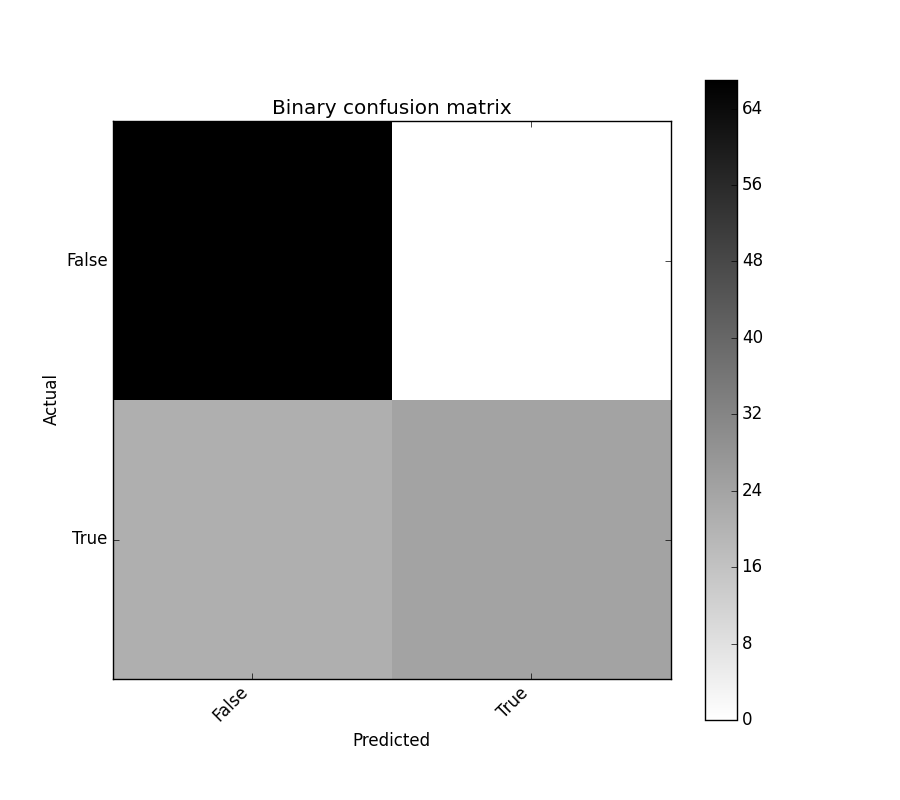
binary_confusion_matrix
Matplotlib plot of a normalized binary confusion matrix¶
binary_confusion_matrix.plot(normalized=True)
plt.show()
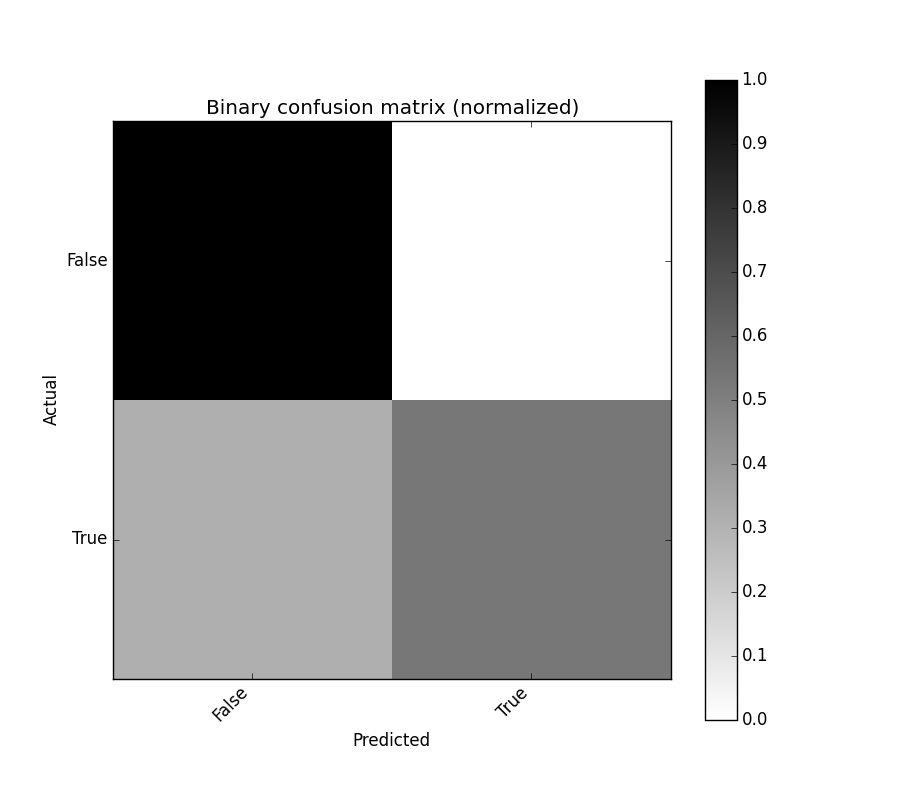
binary_confusion_matrix_norm
Seaborn plot of a binary confusion matrix (ToDo)¶
binary_confusion_matrix.plot(backend='seaborn')
Confusion matrix and class statistics¶
Overall statistics and class statistics of confusion matrix can be easily displayed.
y_true = [600, 200, 200, 200, 200, 200, 200, 200, 500, 500, 500, 200, 200, 200, 200, 200, 200, 200, 200, 200]
y_pred = [100, 200, 200, 100, 100, 200, 200, 200, 100, 200, 500, 100, 100, 100, 100, 100, 100, 100, 500, 200]
cm = ConfusionMatrix(y_true, y_pred)
cm.print_stats()
You should get:
Confusion Matrix:
Classes 100 200 500 600 __all__
Actual
100 0 0 0 0 0
200 9 6 1 0 16
500 1 1 1 0 3
600 1 0 0 0 1
__all__ 11 7 2 0 20
Overall Statistics:
Accuracy: 0.35
95% CI: (0.1539092047845412, 0.59218853453282805)
No Information Rate: ToDo
P-Value [Acc > NIR]: 0.978585644357
Kappa: 0.0780141843972
Mcnemar's Test P-Value: ToDo
Class Statistics:
Classes 100 200 500 600
Population 20 20 20 20
Condition positive 0 16 3 1
Condition negative 20 4 17 19
Test outcome positive 11 7 2 0
Test outcome negative 9 13 18 20
TP: True Positive 0 6 1 0
TN: True Negative 9 3 16 19
FP: False Positive 11 1 1 0
FN: False Negative 0 10 2 1
TPR: Sensivity NaN 0.375 0.3333333 0
TNR=SPC: Specificity 0.45 0.75 0.9411765 1
PPV: Pos Pred Value = Precision 0 0.8571429 0.5 NaN
NPV: Neg Pred Value 1 0.2307692 0.8888889 0.95
FPR: False-out 0.55 0.25 0.05882353 0
FDR: False Discovery Rate 1 0.1428571 0.5 NaN
FNR: Miss Rate NaN 0.625 0.6666667 1
ACC: Accuracy 0.45 0.45 0.85 0.95
F1 score 0 0.5217391 0.4 0
MCC: Matthews correlation coefficient NaN 0.1048285 0.326732 NaN
Informedness NaN 0.125 0.2745098 0
Markedness 0 0.08791209 0.3888889 NaN
Prevalence 0 0.8 0.15 0.05
LR+: Positive likelihood ratio NaN 1.5 5.666667 NaN
LR-: Negative likelihood ratio NaN 0.8333333 0.7083333 1
DOR: Diagnostic odds ratio NaN 1.8 8 NaN
FOR: False omission rate 0 0.7692308 0.1111111 0.05
Statistics are also available as an OrderedDict using:
cm.stats()